The WebApp is sponsored by: 

Improving the diagnostic accuracy of Alagille Syndrome genetic diagnosis using artificial intelligence
Catalina Jaramillo1, Barry Moore2,3, Stephen Guthery1,3.
1Department of Pediatrics, Division of Pediatric Gastroenterology, Hepatology and Nutrition, University of Utah/ Primary Children's Hospital, Salt Lake City , UT, United States; 2Fabric Genomics, Oakland, CA, United States; 3Department of Human Genetics, University of Utah, Salt Lake City , UT, United States
Introduction: Alagille Syndrome (ALGS) is an autosomal dominant disorder ALGS caused by mutations in JAG1 and NOTCH2, many of which are classified as variants of uncertain significance (VUS), highlighting the need for improved diagnostic accuracy. This study evaluates in silico artificial intelligence (AI) methods in the genetic diagnosis of ALGS.
Methods: GEM is an AI-based variant prioritization tool that scores and ranks potentially damaged genes, enabling rapid identification of disease-causing variants in genome sequences. A GEM score of ≥1 can identify >90% of all true-positive damaging variants. We generated a simulated dataset using whole-genome sequences from the International Genome Sample Resource. Known ALGS-causing mutations were informatically spiked into a set of case genomes, while unspiked control genomes were matched on genetic ancestry. Human Phenotype Ontology (HPO) terms were extracted from the electronic medical record (EMR) using a natural language platform to enhance GEM prioritization. GEM scores were compared between cases and controls.
Results: GEM scores were generated for 253 JAG1 and ›NOTCH2 mutations and 504 controls. Scores were significantly higher in cases compared to controls for JAG1 (1.69 vs -1.21, P= <0.01) and NOTCH2 (1.28 vs -1.41, P=<0.01). Nearly all spiked-in genes had scores >0, indicating GEM’s ability to distinguish pathogenic mutations from normal genomic variation. 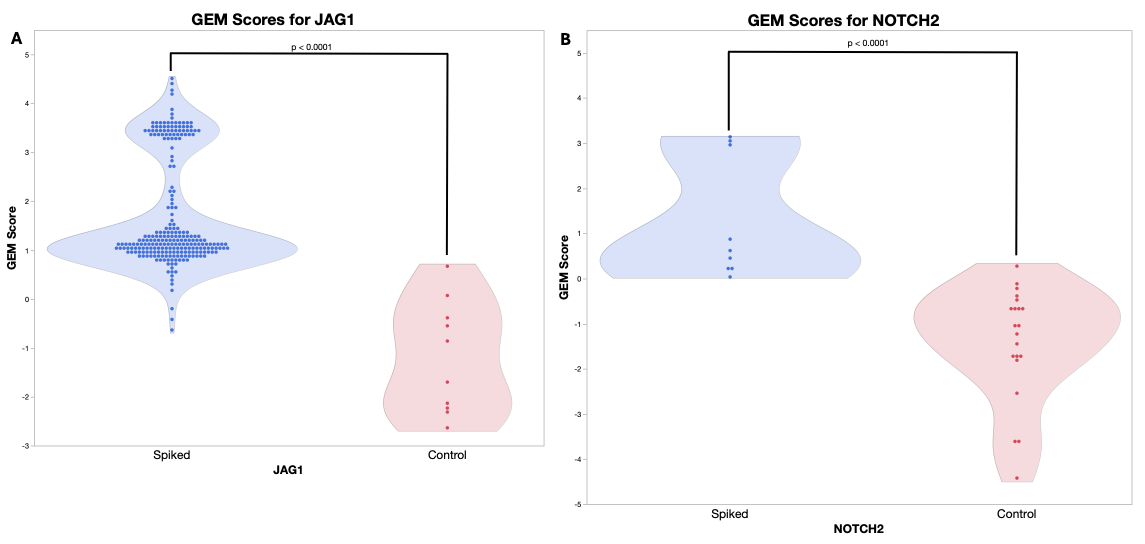
Conclusion: AI-based variant prioritization for JAG1 and NOTCH2 mutations improves diagnostic accuracy in ALGS. HPO term EMR extraction is a key a priori factor to optimize AI-driven ALGS molecular diagnostics. GEM is a promising tool to reduce VUS in ALGS.
This work was supported by the University of Utah “CTSI Partner Mentored Career Development Scholar Award”; and The University of Utah “Center for Genomic Medicine Pilot Award”..
Email: info@splitmeeting.org
If you have any questions during the meeting, please go to the registration desk. Our emails will be monitored sporadically.
REGISTRATION DESK OPENING TIMES